When the cellranger-arc mkfastq or cellranger-arc count pipelines fail, they will automatically generate a "debug tarball" that contains the logs and metadata generated by the pipestance leading up to failure. This file, named sample_id.mri.tgz, can be e-mailed to the 10x Genomics support team to help resolve any issues with using Cell Ranger ARC. You may also use the cellranger-arc upload command to send the tarball to 10x Genomics. Run this code after replacing your@email.edu with your email:
$ cellranger-arc upload your@email.edu sample_id.mri.tgz
If you wish to troubleshoot a pipeline failure yourself, it is important to identify if it is a preflight failure, an in-flight failure, or an alert.
The remainder of this guide uses the term pipestance to refer to a specific instance of a pipeline running.
Preflight failures are the most common and are the result of invalid input data or runtime parameters. Because they occur before the pipeline runs, there will be no pipeline output and the error is reported directly to your terminal.
Common preflight failures include failing to install bcl2fastq. cellranger-arc mkfastq will generate the following error if Illumina's bcl2fastq software is not installed:
[error] On machine: workstation.university.edu, bcl2fastq or configureBclToFastq.pl not found on PATH.
In-flight failures are generally the result of factors external to the pipeline such as running out of system memory or disk space. Since different stages may fail in different ways, the specific error messages will vary widely.
There are a few important files that are saved to your pipeline output directory which, by default, is named according to the flow cell serial number for cellranger-arc mkfastq (e.g., HAWT7ADXX) and your --id name for cellranger-arc count.
- The pipeline execution log that is output to your terminal during pipeline execution is also saved to
output_dir/_log. - Stages that experience a hard failure generate an
_errorsfile containing the precise error that caused a stage to halt. You can view these error logs, if they exist, usingfind output_dir -name _errors | xargs cat - Each stage also logs its stdout and stderr streams to
_stderrand_stdoutfiles. These logs can be listed usingfind output_dir -name _stderrand may contain elucidating error messages in stages that execute third-party applications such as STAR.
A more detailed description of the pipeline output directory and its contents is provided in the Pipestance Structure page.
If you are unable to diagnose a failure yourself, you can always contact the 10x Genomics software support team for help.
Once you have determined the reason for failure and are ready to continue running the pipeline, you can typically issue the same cellranger-arc command to continue execution of the pipestance from the stage that originally failed.
When running cellranger-arc mkfastq or cellranger-arc count, it will detect if its intended output directory already exists. If it does, this existing pipeline output directory will be treated as an incomplete pipestance and resume execution. This feature allows pipelines to be stopped and resumed with great flexibility, but it can also result in errors such as:
RuntimeError: /home/jdoe/runs/sample345 is not a pipestance directory
which indicates that you specified a --id that corresponds to an existing directory that was not created by Cell Ranger ARC.
The following error:
RuntimeError: pipestance 'HAWT7ADXX' already exists and is locked by another Martian instance. If you are sure no other Martian instance is running, delete the _lock file in /home/jdoe/runs/HAWT7ADXX and start Martian again.
indicates that you may already have a copy of cellranger-arc running that is using the same output directory. If you are sure that there is no pipestance running in the given output directory, you can either remove that output directory entirely (mv HAWT7ADXX HAWT7ADXX.old) to restart the pipestance from the beginning, or you can remove the pipestance's lock file (rm HAWT7ADXX/_lock) and re-run the cellranger-arc command to resume pipeline execution.
If you encounter the following error when attempting to resume a pipestance:
RuntimeError: pipestance 'sample345' already exists with different invocation file /home/jdoe/runs/sample345/_invocation
you are attempting to resume a pipestance using command-line arguments that are different from those used to first run it. You can view the parameters input to the existing pipeline by examining the _log file located in the output directory (e.g., head -n20 /home/jdoe/runs/sample345/_log)
Alerts are generally the result of factors inherent in library preparation and sequencing instead of software. Abnormal data (including common short-read sequencing metrics and Chromium Single Cell Multiome ATAC + Gene Expression-specific statistics) are raised in the form of alerts that are printed in the output web_summary.html file. Alerts do not affect the operation of the pipeline, but they do highlight potential causes for abnormal or missing data.
Alerts come in two severity levels:
WARNalerts indicate that some parameter is suboptimal, but there may still be useful data in the pipeline output.ERRORalerts indicate a major issue, and there are unlikely to be usable results in the output.
For example, running cellranger-arc count at lower than recommended sequencing depth will raise the following alerts:
A single WARN and ERROR alert is not indicative of a lost sequencing run but is simply highlighting a reason why the results might be less than ideal.
However, a mis-match between the reference genome and gtf will have an extensive impact on the analysis and raise several ERROR level alerts. For example, running cellranger-arc count with hg19 genome fasta sequences and a GRCh38 gtf will raise the following alerts:
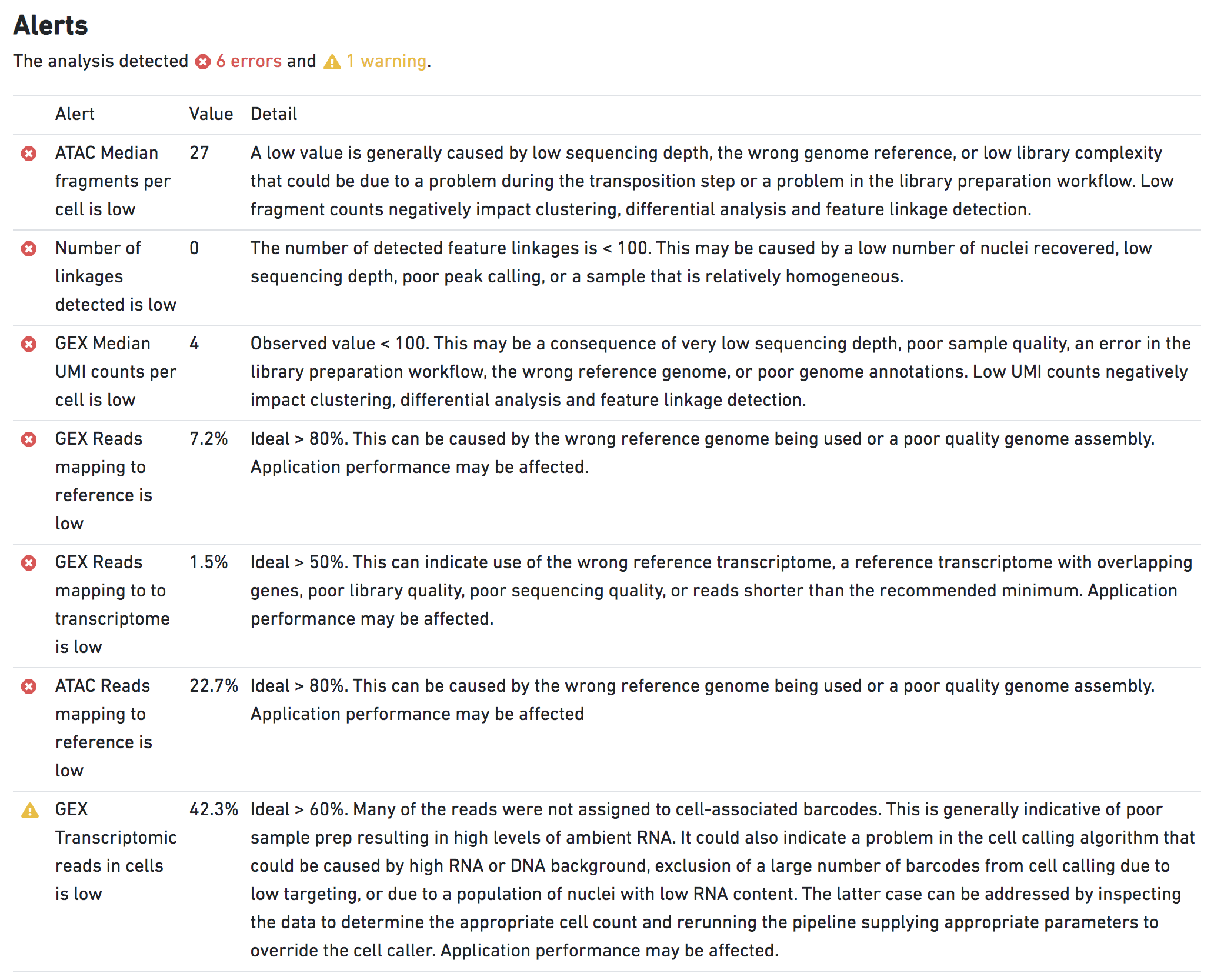
The presence of these ERROR alerts indicate that results output by this pipestance are likely dubious.